Relocation
Due to the demolition of Cowley Hall and construction of Prairie Springs Science Center Phase II, the Biology Department will be temporarily relocated to 212 Cartwright Center. We apologize for any inconvenience.
Undergraduate programs
Biology
Undergrad major Undergrad minor Teacher licenseMajor in biology! If you want to prepare for a career in healthcare, learn about organisms worldwide or help cure diseases, biology may be right for you.
Areas of study
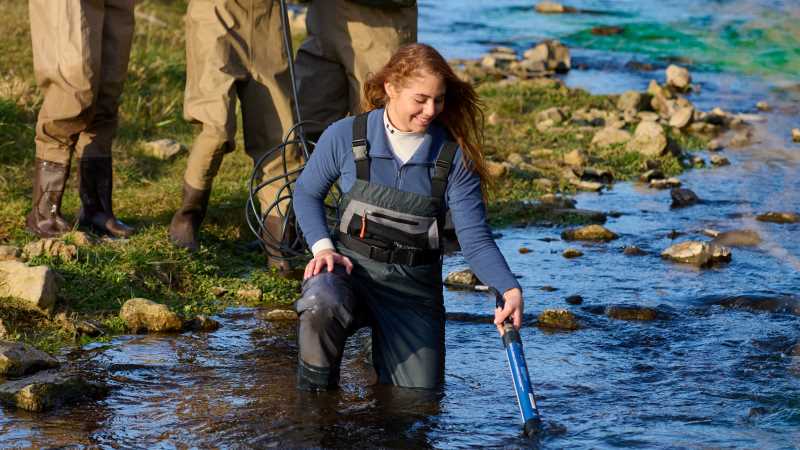
Aquatic Science Concentration
UWL offers an aquatic science concentration within the biology major to prepare students for exciting and challenging careers in the study and management of aquatic resources such as wetlands, streams, lakes, rivers, springs and groundwater.
Undergrad major View a sample plan for Aquatic Science Catalogfor Aquatic ScienceBiomedical Science Concentration
Biomedical science concentration coursework focuses on human and mammalian biology. An excellent choice for pre-med, pre-vet and pre-health professions students, the concentration includes a highly regarded, two-semester human anatomy and physiology series and additional chemistry classes.
Undergrad major View a sample plan for Biomedical Science Catalogfor Biomedical ScienceConservation Biology
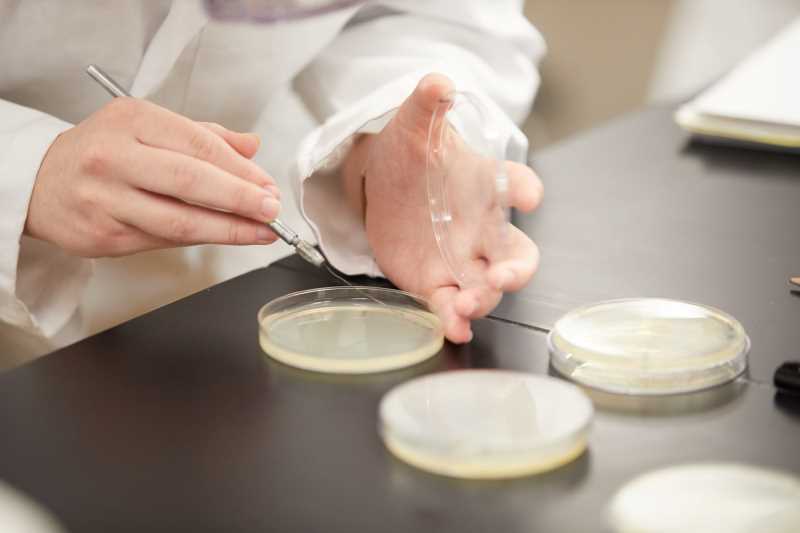
Undergrad majorMolecular Genetics & Cell Biology Concentration
Molecular Genetics and Cellular Biology concentration coursework focuses on understanding living processes at a molecular level. Scientists are making exciting biological discoveries in these fields today whether identifying genes responsible for cancer or re-writing the genetic code of living cells.
Undergrad major View a sample plan for Molecular Genetics & Cell Biology Catalogfor Molecular Genetics & Cell BiologyPlant & Fungal Biology Concentration
Plant & Fungal Biology concentration coursework will help you learn to identify plants and fungi, study their evolution, and understand their physiology. Students can also conduct research with faculty who study plants and fungi. UWL offers one of the only plant and fungal biology degrees in the U.S.
Undergrad major View a sample plan for Plant & Fungal Biology Catalogfor Plant & Fungal BiologyScience Education
Completion of the Biology: Science Education Concentration Program and associated benchmark assessments will lead to endorsement a Middle and High School Science, grades 4-12 (2600), Wisconsin teaching license.
Undergrad major Teacher license View a sample plan for Science Education Catalogfor Science EducationZoology and Animal Physiology
Undergrad majorBiology & Physical Therapy
UWL offers a dual degree program for qualified students, combining an undergraduate Biology degree with a graduate Physical Therapy degree.
Food & Nutrition Sciences
Undergrad majorMajor in food and nutrition sciences. Gain a broad understanding and skillset related to food science, food safety, food systems and nutrition.
Related programs from other departments
Graduate programs
Biology
Graduate degreeAdvance your expertise in biology through hands-on research, individualized mentorship, and advanced scientific training with an M.S. in Biology.
Areas of study